Overview of datasets
We used four datasets. Two datasets are video datasets involving different clinical populations and sensor settings: the dementia video dataset comprises videos recorded with a fixed camera from the front or back collected from patients with dementia and MCI as well as older CU adults, and the CP video dataset comprises videos recorded from the side with a horizontally rotating camera collected from children and younger adults with CP. For the ground-truth labels, both datasets include gait parameters derived from optical motion-capture data as well as clinical measures that can be used for downstream tasks. Since videos and motion-capture data in the CP dataset were not collected simultaneously, model performance in terms of estimating gait parameters is assumed lower than that for the dementia dataset, in which videos and motion-capture data were collected simultaneously, due to intra-individual, trail-to-trial variability13. The other two datasets are accelerometer datasets involving different clinical populations and sensor settings: the dementia accelerometer dataset comprises accelerometer data from the waists or ankles collected from patients with dementia and MCI as well as older CU adults, and the PD accelerometer dataset comprises accelerometer data on the chest or thighs collected from patients with PD and healthy controls of younger and older adults. Both accelerometer datasets have stopwatch-based gait-speed data as a ground-truth label for the gait-parameter-estimation task. A subset of the dementia accelerometer dataset can be supplementarily analyzed with the ground-truth gait parameters derived from optical motion-capture data acquired on different days. The dementia video and accelerometer datasets are our in-house datasets curated in Japan for this study, while the CP and PD datasets are third-party public datasets curated in USA13,56. The study was conducted with the approval of the Ethics Committee, University of Tsukuba Hospital (H29-065, R1-137, and R1-168), and it followed the ethical code for research with humans as stated in the Declaration of Helsinki.
Dementia video dataset
We analyzed a total of 7611 gait videos from 160 unique older adults with dementia (n = 35) and MCI (n = 63) as well as CU individuals (n = 52) and those without formal diagnosis (n = 10; see Supplementary Method 3 for the diagnostic details). The average participant age was 73.5 years (standard deviation (SD): 5.7), with 13.1 years of education (SD: 2.5). Their average height was 1.58 m (SD: 0.09). The participant demographics (self-reported sex, age, years of education, and height) as well as their cognitive, clinical, and gait profiles for each diagnostic group are presented in Supplementary Table 1. All data were collected at University of Tsukuba Hospital, Japan, between 2018 and 2019 (baseline) and between 2021 and 2022 (3-year follow-up). All participants provided written informed consent.
During a lab visit at the baseline, gait trials were video-recorded using fixed cameras from the front and back. The participants completed an average of 12.2 gait trials (SD: 2.1). In each trial, they were asked to walk 9.5 m at their usual pace. The walkway was a flat space 12 m long × 3 m wide without obstacles. The 9.5-m path started at 2.5 m from a short edge of the walkway and ended when crossing the opposite edge. Four commodity webcams were fixed approximately 1.25 m high at each of the four corners of the walkway, such that their angles of view looked approximately towards the center of the walkway. Each gait was recorded using the four cameras, which resulted in four videos per trial. Videos were stored in an MP4 format at a resolution of 720 × 1280 or 1080 × 1920 pixels for width and height and a rate of 30.0 frames per second.
For each video frame, we extracted 2D image-plane coordinates of body keypoints by using the 26-keypoint RTMPose model57, a state-of-the-art keypoint-extraction model at the time of analysis, which resulted in a time series of (x,y)-coordinates of keypoints along with a confidence score of 0 to 1 for each keypoint for each frame. After removing keypoints with a confidence score of less than 0.3, videos were automatically excluded from the analysis if (i) it was shorter than 60 frames (2 s) or (ii) did not contain a gait of at least four steps estimated by the number of peaks in the y-coordinate of ankle keypoints. Each time series was then truncated to exclude the first and last two steps or more to discard those affected by increased and decreased speed. Errors in keypoint detection, e.g., confusion of different keypoints, were corrected manually for each frame by re-labeling mis-identified keypoints or by excluding erroneous keypoints from the analysis. This procedure yielded a total of 7611 time-series data samples of an average length of 157 frames (SD: 42 frames) or of 5.2 s (SD: 1.4 s; Supplementary Fig. 5a for the full distribution).
Ground-truth gait parameters were computed using optical 3D motion-capture data concurrently recorded in the same gait trials. We used the eight-camera OptiTrack Flex 13 motion-capture system (NaturalPoint, Inc., Corvallis, OR, USA), sampled at 120 Hz with 50 reflective markers placed on pre-defined anatomical landmarks according to the manufacturer’s instructions. The data were semi-manually post-processed to correct marker tracking errors. To discard the increase and decrease speed effect, the first two steps and last 2.5 m were excluded from analyses. The gait parameters were computed on the basis of the 3D body kinematics measured using the motion-capture system, where the gait speed was calculated using the trajectory of the pelvis marker on the waist back, and the step measures were calculated using the trajectories of the heel markers. Specifically, the step measures were calculated by identifying stationary time-segments on the ground in the following steps. First, the position trajectories of the heel markers were interpolated using a piecewise cubic polynomial interpolation algorithm and low-pass filtered with a cut-off frequency of 10 Hz, which was used for calculating the velocity. We then identified stationary time-segments on the basis of the velocity of the heel markers and obtained timing of heel strikes and positions of heels at stationary phase. All results were plotted and visually confirmed. Finally, step time was calculated as the time interval between heel strikes, and step length and step width were calculated using the distance relative to the anteroposterior and mediolateral axes between the heel positions at successive stationary phases, respectively. Due to a number of missing markers in the motion-capture data, which precluded post-processing and fair calculation of gait parameters, we did not calculate ground-truth gait parameters for motion-capture data with initial marker-missing rates of approximately 20% or more and calculated them for three gait trials per participant if available. Consequently, ground-truth gait parameters were available for 1423 out of the 7611 videos. Regarding the downstream tasks, a total of 7122 videos from 150 participants with the baseline diagnosis were included in the analysis. Specifically, 7122, 4006, and 4373 videos from 150, 87, and 89 participants were used for the cross-sectional multi-class and binary-classification tasks on the dementia diagnostic status as well as the longitudinal-prediction task on cognitive decline, respectively. The cognitive decliners were defined as those with the 3-year follow-up Mini-Mental State Examination58 score of two or more points below the baseline score, resulting in 27 of the 89 participants (1292 of the 4373 videos) identified as a cognitive decliner. We also considered the following auxiliary variables for model analyses: the height for the gait-parameter estimation, assuming that it is essential as the basis for estimating spatial parameters, and the age, years of education, and height for the downstream tasks.
In this dataset, the participants underwent muscle-activity measurement simultaneously with video recording. To record muscle activity during gaits, we used 16 wireless surface EMG sensors sampled at 2000 Hz (Trigno Wireless EMG System, Delsys, Inc., Natick, MA, USA). These sensors were directly attached to the skin using double-sided adhesive tape, following skin preparation with alcohol wipes. Sensors were placed following SENIAM guidelines ( with bilateral placement over the following lower limb muscles: rectus femoris, biceps femoris, tibialis anterior, medial gastrocnemius, vastus medialis, peroneus longus, soleus, and medius gluteus muscle. As EMG data for the medius gluteus muscle could be collected only from a subset of the participants, these data were excluded from the present analysis. We further excluded data from the analysis if they met either of the following criteria: (i) missing values due to technical issues such as battery failure or transmission errors, or (ii) lack of substantial change in EMG values between stationary standing and walking, defined as the average EMG value during walking smaller than the mean+5 SD during stationary standing, which could be attributed to inadequate sensor attachment or skin sweat. This procedure yielded a total of 3566 video data samples paired with EMG data (average length: 5.2 s, SD: 1.4 s) from 150 unique participants consisting of patients with dementia (n = 32) and MCI (n = 58) as well as CU individuals (n = 50) and those without formal diagnosis (n = 10).
Cerebral-palsy video dataset
We analyzed a total of 1636 gait videos from 968 unique patients, mainly children, with CP diagnosis. The average age of the patients included in the analysis was 12.6 years (SD: 6.1). Their average height, mass, and gait deviation indices (GDIs) were 1.41 m (SD: 0.19), 40.2 kg (SD: 17.1), and 76.5 (SD: 10.9), respectively. The patient demographics (age, height, and mass) as well as clinical and gait profiles are detailed in Supplementary Table 2. The participants underwent an average of 1.2 gait visits (SD: 0.5), where each visit comprised one gait trial for video recording and another for ground-truth recording. All data were collected during their clinical gait visits conducted in USA. See the original article13 for details of the dataset.
Videos were recorded during gait trials using an unfixed, rotating camera from the sagittal view. In each trial, the patients were asked to walk back and forth along a 10-m path three to five times. Each gait was video-recorded using a single commodity camera placed approximately 3 to 4 m from the line of walking, where the camera was manually rotated around the vertical axis to follow the patient. Videos were stored in an MP4 format at a resolution of 640×480 pixels for width and height and a rate of 29.97 frames per second.
For each video frame, the time-series of (x,y)-coordinates of body keypoints were provided along with confidence scores, where keypoints were detected using the 25-keypoint OpenPose model59. As this is a third-party dataset provided solely as pre-extracted keypoint data, the model used for keypoint extraction was different from that used for the dementia video dataset. However, this methodological difference does not impact the interpretation of our investigation, as our objective was to demonstrate the utility of the proposed approach across datasets, rather than conduct a comparative performance analysis between the datasets. Specifically, we first split each time-series data sample into multiple segments of a straight walk by dividing the data at the turn timings. After removing keypoints with a confidence score of less than 0.3, segments were automatically excluded from the analysis if (i) they were shorter than 60 frames (2 s), (ii) any keypoint necessary for the analysis was missed for two or more consecutive frames, or (iii) the patient stopped in the middle of walking. A stop in the walk was detected by keypoint movement of ankles smaller than 0.05 in the image-plane coordinates normalized by the length between neck and middle hip. This procedure yielded a total of 1636 time-series data samples of an average length of 188 frames (SD: 108 frames) or of 6.3 s (SD: 3.6 s; Supplementary Fig. 5b for the full distribution).
Ground-truth gait parameters were provided for all body keypoint data on the basis of optical 3D motion-capture data separately recorded in the same lab visit. Specifically, the 12-camera Vicon MX motion-capture system (Vicon Motion Systems Ltd, Oxford, UK), sampled at 120 Hz, was used. The data were semi-manually post-processed to fill missing marker measurements then used to compute gait parameters of interest from 3D joint kinematics. Although the gait parameters were provided for each side in the original dataset, we used the averages of left and right values in our analysis for consistency across datasets. For the downstream task, the GMFCS level, a widely used five-level clinical grading system to assess the self-initiated movement abilities of patients with CP60, was rated by a physical therapist. The patients were rated as level I (lowest severity; n = 277), level II (n = 245), level III (n = 128), and level IV (n = 3), with no patients rated as level V (highest severity), yielding a total of 911 videos included in the analysis. We excluded level IV from our analysis due to the limited number of data samples, as it would make it impractical to conduct stratified cross-validation, as described in the subsequent section. For model analyses, we also considered the following auxiliary variables: the height for the gait-parameter estimation and height, mass, age, and GDI for the downstream task.
Dementia accelerometer dataset
We analyzed a total of 2489 wearable accelerometer data samples captured during gait trials from 164 unique older adults. The average participant age was 73.1 years (SD: 7.0) and the group included patients with baseline clinical diagnosis of dementia (n = 19) or MCI (n = 57) as well as CU individuals (n = 60). The participants included those diagnosed in routine clinical practice, for whom the diagnostic criteria may differ from those for other participants. The participant demographics (self-reported sex, age, and years of education) as well as their clinical and gait profiles for each diagnostic group are presented in Supplementary Table 3. All data were collected during lab visits or through a paid cognitive health checkup service in clinical practice (see Supplementary Method 1 for details) at the University of Tsukuba Hospital, Japan, between 2018 and 2023. All participants provided written informed consent.
Accelerometer data were recorded during gait trials using tri-axial accelerometers mounted on participants’ waists and both ankles. We used GENEActiv sensors (Activinsights Ltd, Kimbolton, UK), sampled at 100 Hz. Each ankle sensor was firmly attached to the outside of each ankle, just above the lateral malleolus, using Velcro bands with the sensor’s y-axis pointing downwards. The waist sensor was firmly attached to the lower back approximately on the L5 position, a commonly used sensor placement for approximating the center of mass, using surgical tape with the sensor’s y-axis pointing downwards.
In each gait trial, the participants were asked to walk flat corridors without any obstacles. We used three different distances of 4, 5, and 7 m for gait measurement, excluding acceleration and deceleration paths of 2 m or more, where the distance varied depending on the availability of the facility. The elapsed time necessary to walk the path was manually measured using a stopwatch for recording timestamps at timings when the participant crossed the start and end lines for the measurement. The accelerometer data between the start and end timestamps were cropped for each trial and used for the analysis. In summary, each participant completed an average of 4.0 gait trials (SD: 0.5). We excluded data from the analysis for (i) inconsistency in sensor orientation and (ii) errors in timestamp recording. This procedure resulted in a total of 2489 time-series data samples (803 for waist and 1686 for ankles) of an average length of 486 frames (SD: 123 frames) or of 4.9 s (SD: 1.2 s; Supplementary Fig. 5c for the full distribution), along with the stopwatch-based ground-truth gait-speed data.
For this dataset, 89 of the 164 participants overlapped with those included in the dementia video dataset, whereas all accelerometer data were captured in different visits. The accelerometer data for 68 of the 89 participants were collected at least one year after video- and motion-capture data collection. For the other 21 participants, some accelerometer data were collected within a shorter period with the video and motion-capture data (mean interval: 7.2 days, SD: 7.6 days; Supplementary Table 4), yielding 401 accelerometer data samples labeled with optical-motion-capture-based ground-truth gait parameters.
Parkinson’s disease accelerometer dataset
We analyzed a total of 382 wearable accelerometer data samples captured during gait trials from 32 unique individuals with PD (PD; n = 16) and controls (n = 16). The average age of the participants included in the analysis was 64.5 years (SD: 10.5). The details of the participant demographics (sex and age) as well as their clinical and gait profiles for each diagnostic group are presented in Supplementary Table 5. The gait data were collected during in-clinic assessments conducted in USA. See the original article56 for details for this dataset.
The accelerometer data included in our analysis were recorded during gait trials using tri-axial accelerometers mounted on participants’ chests and each anterior thigh. BioStamp RC sensors (MC 10 Inc., Lexington, MA) were used at a sample frequency of 31.25 Hz. The chest sensor was attached with the sensor’s y-axis pointing downwards. Each thigh sensor was attached with the sensor’s x-axis pointing upwards.
As the gait trials, the participants underwent 10-m walk tests as part of standard in-clinic assessments for PD. The elapsed time necessary to walk the path was measured using manually recorded timestamps at the start and end of the test, which is equivalent to measurement with a stopwatch. The accelerometer data between the start and end timestamps were cropped for each trial and used for the analysis. In summary, each participant with PD completed an average of 2.8 walk test trials (SD: 0.4) in the ON state and 2.9 trials (SD: 0.5) in the OFF state. The control participants completed 3.1 trials (SD: 0.2). We excluded data from the analysis for (i) inconsistency in sensor orientation and (ii) errors in timestamp recording. This procedure resulted in a total of 382 time-series data samples (131 for chest and 251 for thighs) of an average of 148 frames (SD: 33 frames) or of 4.8 s (SD: 1.1 s; Supplementary Fig. 5d for the full distribution), along with the stopwatch-based ground-truth gait-speed data.
Overview of proposed approach
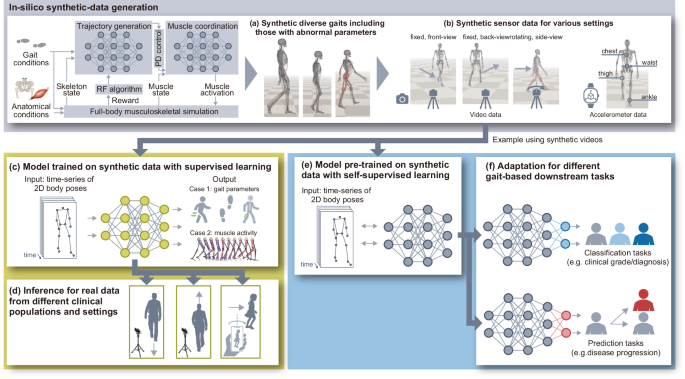
Our proposed approach, illustrated in Fig. 1, consists of four main steps: (i) in-silico generation of synthetic gaits with a deep generative model learned in the musculoskeletal simulation (Fig. 1a); (ii) in-silico generation of synthetic sensor data derived from the synthetic gaits (Fig. 1b); and (iii) model training on synthetic data with supervised learning (Fig. 1c) followed by (iv) inference with the model on real data (Fig. 1d), or (iii’) model pre-training on synthetic data with self-supervised learning (Fig. 1e) followed by (iv’) fine-tuning of and inference with the model on real data for each downstream task (Fig. 1f). We use a single-camera video as an example to describe the approach.
Regarding the in-silico generation of synthetic gaits (i.e., the first step of our approach), we use Generative GaitNet33 that learns muscle control of human gaits in accordance with the desired anatomical and gait conditions through deep reinforcement learning and a full-body musculoskeletal model in physics-based simulation. We generate diverse synthetic gaits that represent a broad spectrum of gait and anatomical conditions by continuously modifying simulation parameters, including target gait parameters (e.g., stride length and cadence), body properties (e.g., body size and proportion of body parts), and muscle-deficit characteristics (e.g., weakness and contracture). Atypical synthetic gaits with abnormal parameters are included to increase the diversity of the synthetic data. In the second step, we simulate sensor data during these simulated gaits. For video data, we project three-dimensional (3D) body poses into a 2D plane, then obtain the time series of the 2D coordinates of body poses, equivalent to the data produced using computer-vision-based pose-estimation algorithms applied to real gait videos. To improve the model’s generalizability across different sensor settings, we use the domain-randomization technique to generate synthetic sensor data, i.e., 2D body poses, for different camera positions, postures, and movements (i.e., fixed or rotating to follow the person). The synthetic data are used in the following two ways. The first involves using the time-series of 2D body poses and gait parameters derived from synthetic gaits (i.e., gait speed, step time, and step length) as input and output variables, respectively, for model training with supervised learning. The models trained solely on synthetic data are then applied to real data for gait-parameter estimation from 2D, single-camera gait videos. The second way involves using the time-series of 2D body poses as pre-training data for learning general-purpose representations through self-supervised learning. The pre-trained model is then fine-tuned with real data for each downstream task and tested with separate real data.
In-silico synthetic data generation
We used the pre-trained Generative GaitNet with a full-body musculoskeletal model33 and generated synthetic gaits with different parameter sets related to gait and anatomical conditions to emulate human gaits of adults and children, including pathological gaits, included in the datasets used in the analysis. The musculoskeletal body model has 50 degrees of freedom and 304 Hill-type muscle models, and its dynamics was simulated with 480 Hz on the DART physics engine61. The reference skeletal body model represents an individual of 168.7 cm in height and 72.9 kg in weight, and the body size and proportions of each body part of the model, as well as gait and muscle conditions, can be arbitrarily modified in the simulation by specifying the parameters. We modified a total of 132 continuous parameters consisting of 6 parameters for body properties, 2 parameters for gait conditions, 62 parameters for muscle contracture of lower limbs, and 62 parameters for muscle weakness of lower limbs. All parameters are scale factors with respect to the reference model. The body parameters are the scale factors of the whole-body size, as well as the proportion of each body part for the head and four lower limbs. The parameters for gait conditions are for specifying target stride length and cadence. The parameters for muscle weakness and contracture are the scaling factors of the maximal isometric force and optimal length for each muscle, respectively. We separately prepared parameter sets for adult- and children-like bodies as well as for normal and abnormal muscle conditions. While the scale factors of body size for adults were randomly generated with a uniform distribution ranging from 0.85 to 1.1, those for children were randomly generated with a normal distribution, N(0.75, 0.1) and clipped to fall within [0.5, 1]. While parameters for the normal muscle conditions were set to 1, those for abnormal muscle conditions each were randomly generated with a normal distribution, N(1, 0.05) and clipped to fall within [0.7, 1]. Other parameters were commonly generated as follows. The scaling factors for proportions of each body part were randomly generated with a uniform distribution ranging from 0.95 to 1.05, and the scaling factors for stride length and cadence were randomly generated with a uniform distribution ranging from 0.6 to 1.4 and from 0.8 to 1.4, respectively. We randomly generated 3000 parameter sets each for synthetic gaits of adults and children with normal and abnormal parameters of muscles, yielding a total of 12,000 parameter sets for synthetic gaits used in this study. We ran the simulation at 480 Hz with each parameter set and used data segments of a 10-m walk from the 5-m point in simulation runs where the musculoskeletal models were able to walk 20 m without falling. The parameters for body size and gait conditions were determined to have a wide range, including values from the datasets used in this study. The parameters for abnormal muscle conditions were determined on the basis of preliminary experiments to avoid too many failed runs where the musculoskeletal models were unable to walk 20 m due to falling. We used a laptop, HP OMEN 15-dh1004TX, with Intel Core i9-10885H CPU 2.40 GHz and a NVIDIA GeForce RTX 2080 SUPER with Max-Q design to run gait simulations. The simulation runs in almost real time. For more details including the learning algorithms of Generative GaitNet, see the original article33.
We then generated synthetic sensor data for videos and accelerometers during the simulated gaits. Regarding the videos, we projected 3D body poses into a 2D image plane assuming no lens distortion, emulating the data format after applying computer-vision-based pose-estimation algorithms into videos. Before the projection, we added random noise following a normal distribution N(0, 0.02) [m] into 3D body keypoints, assuming errors of pose estimation in real data. This value was determined on the basis of the mean per joint position errors of state-of-the-art 3D human-pose-estimation models on walking videos (e.g., around 11 to 25 mm62). The cameras were placed to emulate the camera settings in the datasets used in this study then translated and rotated randomly. To emulate fixed cameras from the front or back in the dementia dataset, one camera was placed 1.5 m to the mediolateral axis and 2.0 m backwards from the start position of the 10-m walk, while another camera was placed 1.5 m to the mediolateral axis and 2.0 m forwards from the end position of the 10-m walk. Both cameras were placed 1.25 m high, and rotated horizontally so that each camera faced the center of the 10-m walkway. Each camera was then translated from the reference position with random offsets each in the mediolateral, anteroposterior, and vertical axes, the values of which were determined with uniform distributions ranging ±0.025, ±0.15, and ±0.025 m, respectively. Finally, each camera was rotated in yaw, roll, and pitch with random degrees determined by uniform distributions ranging ±12.5, ±5.0, and ±5.0, respectively. To emulate horizontally rotating cameras from the side in the CP dataset, the camera was placed 1.0 m high and 3.5 m to the mediolateral axis from the center of the 10-m walkway. The camera was then translated from the reference position with random offsets each in the mediolateral, anteroposterior, and vertical axes, the values of which were determined with uniform distributions ranging ±1.5, ±0.1, and ±0.3 m, respectively. The camera’s initial posture was also rotated randomly in pitch and roll with random degrees determined from uniform distributions ranging ±12.5 and ±5.0, respectively. During the gait sequence, the camera was horizontally rotated to follow the center of mass of the body model with random noise following a normal distribution N(0, 0.5) [degrees]. To emulate accelerometer data in the datasets used in this study, that is, accelerometers attached to the waist, ankle, chest, and thigh, we used the acceleration of the following four body parts in each body coordinate: pelvis, tibia, torso, and femur. These body and gravitational accelerations were described in the coordinate system of each body part, then these data were used as synthetic accelerometer data. Similar to the camera, each coordinate of synthetic accelerometers was rotated randomly in yaw, roll, and pitch with random degrees determined by uniform distributions ranging ±30.0, ±15.0, and ±15.0, respectively. These random values for both synthetic videos and accelerometers were independently calculated for each synthetic sensor data sample.
Preprocessing of time-series sensor data
Both synthetic and real sensor data were converted into the time-series data of F features derived from sensor data with the same preprocessing procedures unless otherwise specified. After the preprocessing, we randomly extracted 30 segments of T-seconds (T is segment length) each, allowing for overlaps, from each time-series data sample of features. These segments served as fixed-length inputs for DNN models. The T was set to 2 and 1 s for the gait-parameter estimation and downstream tasks, respectively, applicable to both video and accelerometer data. The left-right axis was inverted both for videos recorded from the right side and for accelerometers acquired from the body part on the right side to simplify the subsequent training of DNN models.
The body keypoint data derived from each single video was processed in the following five steps: (i) resampling, (ii) interpolation for missing values, (iii) smoothing, (iv) coordinate transformation (scaling and translation), and (v) feature extraction. The body keypoint data were first resampled to 30 Hz, a standard sampling rate of video data, yielding 60- and 30-frame segments as DNN model inputs for gait-parameter estimation and downstream tasks, respectively. Because the real data used in this study were recorded at about 30 Hz, only synthetic data were down-sampled from 480 to 30 Hz. After imputing missing data using linear interpolation, we then applied smoothing to each time-series data sample using a 1D unit-variance Gaussian filter. In the fourth step, positions of body keypoints in each frame were divided by length \(l\) and translated so that the center of the hip is at the origin of the 2D coordinates, where \(l\) was calculated in each frame using the average of the Euclidean distance between the right hip and right shoulder and that between the left hip and left shoulder in the 2D image plane. Finally, we extracted F features in each frame and obtained time-series data of F features. For the model for estimating gait parameters, we used the following 15 features: 2D positions of lower-limb keypoints in both sides (hip, knee, and ankle), knee angles of both sides in the 2D coordinate formed by the ankle, knee, and hip keypoints in each side, and the Euclidean distance between the left and right ankle keypoints in the 2D coordinate. For the model for the downstream tasks, to capture whole-body dynamics of human gaits, we added 18 other features consisting of 2D positions of other body-part keypoints (shoulder, elbow, wrist, and toe in both sides as well as neck), yielding a total of 33 features.
Accelerometer data were processed in the following three steps: (i) low-pass filtering, (ii) resampling, and (iii) feature extraction. In the first and second steps, we applied a low-pass filter with a cut-off frequency of 30 Hz then resampled to 50 Hz, yielding 100- and 50-frame segments as DNN model inputs for gait-parameter estimation and downstream tasks, respectively. Thus, synthetic and real data in the dementia dataset were down-sampled from 480 and 100 Hz, respectively, while real data in the PD dataset were up-sampled from 31.25 Hz. We then added the Euclidean norm of acceleration to the raw tri-axial acceleration data for each four-body placement we studied (waist, ankle, chest, and thigh) and finally obtained time-series data of a total of 16 features (4 features × 4 placements). Because we focused on data analysis for single-accelerometer data in this study, accelerometer data acquired from different body placements separately formed inputs by using zero-padding to other feature dimensions (i.e., 12 dimensions). In the supplementary analysis on the subset of the dementia accelerometer dataset with optical motion-capture data, we analyzed waist-worn and ankle-worn accelerometers, yielding eight features. In the model with self-supervised training, we analyzed chest-worn accelerometer data, yielding four features.
DNN-model development and evaluation
For supervised learning, we used the following three models with representative network architectures for analyzing time-series data: fully convolutional network36,37, ResNet36,37, and transformer38. We used each model without any modification of hyperparameters for the network architecture and model training, as implemented in each of the previous study unless otherwise specified (see Supplementary Result 8 and Supplementary Tables 16–19 for model performance with different hyperparameters). Due to the limited size of available real data, we also considered a simpler model architecture with fewer parameters. However, our preliminary analyses on simpler models based on the multi-layer perceptron architecture showed suboptimal performance in estimating gait parameters (see Supplementary Result 9 for details). Consequently, we opted to employ more complex models, as listed above, in the primary analysis. In brief, the fully convolutional network consists of three convolutional blocks and one global average pooling layer fully connected to an output layer. Each convolutional block contains three operations: a 1D convolution followed by batch normalization then rectified linear unit (ReLU) activation. The ResNet consists of three residual blocks and one global average pooling layer fully connected to an output layer. Each residual block contains three convolutional blocks with a linear shortcut linking the output of a residual block to its input. The transformer model consists of four transformer-encoder blocks, one global average pooling layer, and one fully connected layer that is fully connected to an output layer. Each transformer-encoder block contains a multi-head attention layer with dropout, layer normalization, and residual connection, followed by two 1D convolutions with dropout and layer normalization.
For the downstream tasks, we also included the following five models in the comparison: TimesNet39, Non-stationary Transformer40, Informer41, LSTM42, and Mamba43. The first three models show state-of-the-art performance for time-series classification tasks in the public benchmark63, and the last two were added as a representative general sequence model. TimesNet is a convolutional-neural-network-based architecture and designed to capture the temporal patterns derived from different periods with Fast Fourier Transform39. Non-stationary Transformer and Informer are transformer-based architectures40,41. Non-stationary Transformer was proposed as an effective series stationarization architecture that improves the predictive capability of non-stationary series40. Informer is designed to capture long-range dependency between time-series inputs and outputs for the time-series forecasting with a self-attention mechanism based on the query sparsity measurement41. Mamba is a state-space model architecture showing promising performance on various domains including time-series data analysis43. We used these four model architectures as implemented in the Time Series Library64, where the Mamba model, which is originally designed for forecasting tasks, was connected to the classification layers implemented in the TimesNet model with the same parameters. We trained all models for a maximum of 100 epochs with a batch size of 32 or 256, early-stopping by using Adam optimizer, and learning rates of 5 × 10-6 and 5 × 10-4 for video and accelerometer data, respectively.
We carried out self-supervised learning by using the TS2Vec method44 in addition to the above eight models. The TS2Vec method uses a hierarchical contrastive learning framework, in which contrasting is executed for lower-level (frame-level) to higher-level (whole-time-series-level) representations, to learn contextual representations for arbitrary sub-series at multiple levels of time granularity. With this method, positive pairs are defined as representations of the same timestamp from randomly masked views, without any temporal or spatial transformations, while all other pairs are considered negative, enabling the model to effectively learn, we assume, temporal and spatial gait dynamics. We used the same parameters used in the original study44, i.e., a batch size of 8, learning rate of 0.001, and representation dimension of 320. For fine-tuning on downstream tasks, we added a new fully connected layer with L1 regularizer (0.1, 0.01, or 0.001) on top of the frozen pre-trained encoder. This layer ingested the 320-dimensional representation yielded by the encoder and was fully connected to an output for each downstream task. Because our downstream tasks include classification on an imbalanced dataset, we experimented with and without class weights in the loss function, where the class weights were calculated as inverse class frequency. Regarding the other eight models, we first trained the model using synthetic data to forecast the next one second of time-series data (i.e., 30 and 50 frames for video and accelerometer data) then conducted fine-tuning on each downstream task. For fine-tuning on downstream tasks, we replaced the last layer connected to the output layer with one fully connected layer connected to a new output layer for each downstream task and trained this layer while freezing the remaining connections. A validation set was used for selecting the above-mentioned parameters for L1 regularization and class weighting as well as for applying early stopping.
The time-series data derived from videos for these DNN models was standardized in each feature dimension by using the mean and SD of the training set, while the data derived from accelerometers were directly fed into the DNN models. For the regression tasks, the output layer received inputs from previous layers as well as auxiliary variables by using a linear activation function, and the model was optimized using the mean squared error as the loss function. For the classification tasks, the output layer received the same inputs by using a softmax activation function, and the model was optimized using cross entropy as the loss function.
For internal evaluation, we used the subject-wise cross-validation procedure, with which the dataset was split into training, validation, and test sets in the ratio k:1:1, ensuring that each individual’s data were only included in either the training, validation, or test set. For gait-parameter estimation, we used k = 1, 2, 3, 8, 18, and 38, corresponding to training with 33, 50, 60, 80, 90, and 95% of the dataset, respectively. To evaluate the conditions under which the proportion of the training set was even smaller, subsets of the training set were used for actual model training, which were extracted in a subject-wise manner after splitting the dataset into the training, validation, and test sets. For the downstream classification tasks, we used k = 2 and 8 for video and accelerometer data, respectively, with stratified sampling to handle the imbalanced dataset. To mitigate the influence of selection bias and fairly assess the data efficiency, data splitting and subset selection were conducted randomly, and the cross-validation procedure was repeated a minimum of five times for each condition using distinct random seeds. Model outputs were averaged for segments extracted from the same single-sensor data or from the same individual. Model performance was then evaluated using Pearson’s correlation coefficient and mean absolute error for the regression tasks, whereas AUC and balanced accuracy were used for the binary and multi-class classification tasks, respectively. The statistical significance of performance improvement was examined from two-tailed t-tests.
Data efficiency was assessed by determining the amount of real data required to achieve a reference performance. For gait-parameter estimation, this corresponded to the amount of real data needed to train models from scratch to achieve the synthetic-data-trained model’s performance. For downstream tasks, it represented the amount of real data required to fine-tune the synthetic-data-pre-trained model to outperform the best model without pre-training. The required data amount was estimated with linear interpolation between data points, such as estimating performance at 55% of data from models trained on 50 and 60% of data. For gait-parameter estimation, each data point represents one dataset split with varying k, as described above. The video-based downstream tasks were evaluated for 10 to 100% with 10% increments, while the accelerometer-based downstream tasks were evaluated for 20 to 100% with 20% increments, given the smaller datasets.
For comparison, synthetic gait data were also generated using TimeGAN65, a representative GAN-based time-series augmentation technique. Specifically, GANs were trained for 100 epochs using the training dataset for each class in each cross-validation fold during the evaluation of real-data-trained models, and each trained GAN was used for generating synthetic time-series segments to double the size of the training set. Hyperparameters were left as the default values implemented in the Time Series Generative Modeling library66.
Dimension reduction and clustering
We used the UMAP47 and Gaussian mixture model46 to probe the associations between synthetic and real data used in this study. The UMAP was used as dimension reduction to find the low-dimensional subspace where synthetic gaits with normal and abnormal parameters were separated. The input data for the UMAP were vector representations (i.e., embedding vectors) obtained from the TS2Vec model pre-trained on synthetic data (i.e., 320-dimensional vectors), which achieved the highest average ranking across the downstream tasks (Supplementary Table 20). We used the UMAP algorithm with the following parameters: the number of components = 2, metric = “cosine”, minimum distance = 0.99, and number of neighbors = 15. The Gaussian mixture model was used to estimate the probability-density function of the representation vectors for synthetic gaits in the 2D UMAP subspace. When the UMAP was repeatedly applied with different parameters, we could robustly identify six clusters corresponding to the combinations of camera settings (fixed-camera/front-view, fixed-camera/back-view, and rotating-camera/side-view) and parameter types (normal and abnormal) for synthetic gaits. Quantitatively, when the Gaussian mixture model was applied with the number of mixture components of 6, about 90% of the synthetic data of each above-mentioned group belonged to the same cluster, which was different from the clusters dominantly belonging to those of different groups. Thus, we used six components for the Gaussian mixture model. Using the estimated probability-density function, we investigated whether the real data in our datasets fall within 95% of the Gaussian distributions for the synthetic data. We repeated this procedure 100 times for statistical analysis, and the statistical significance was examined using one-way analysis of variance with Tukey’s pairwise comparisons. The Fréchet inception distance48 was calculated for each cluster resulting from the Gaussian mixture model, and the mean value was used for the analysis.
Statistics & reproducibility
The statistical analyses of the data and the reproducibility of experiments are detailed in the Nature Portfolio Reporting Summary linked to this article.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link